Your message has been sent.
 CLOSE SIDEBAR
CLOSE SIDEBAR
Antibiotic-Resistance Bacteria in Atlanta's Urban Waterways
Rella Madeline Arena, Emma Allen, Redesh Rai, Dan Thompson, (Faculty Sponsor Dr. Jessica Joyner)
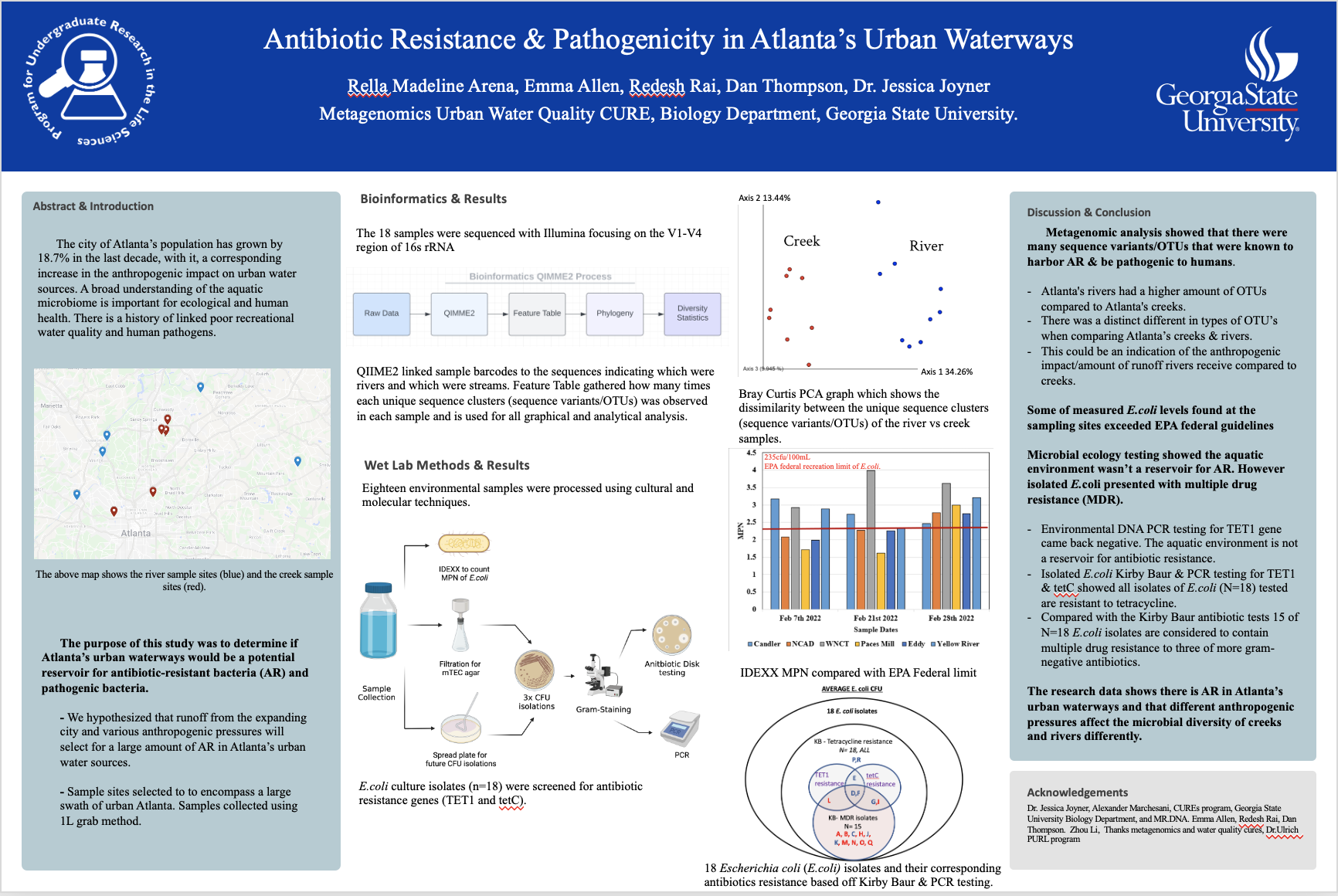
The city of Atlanta’s population has grown by 18.7% in the last decade, with it, a corresponding increase in the anthropogenic impact on urban water sources. A broad understanding of the aquatic microbiome is important for ecological and human health. There is a history of linked poor recreational water quality and human pathogens. The purpose of this study was to determine if Atlanta’s urban waterways would be a potential reservoir for antibiotic-resistant bacteria (AR) and pathogenic bacteria. We hypothesized that runoff from the expanding city and various anthropogenic pressures will select for a large amount of AR in Atlanta’s urban water sources.
Samples were collected throughout Atlanta during summer 2021 & Spring 2022. Microbial diversity was determined with Illumina sequencing of the 16S rRNA gene. We coded a pipeline utilizing QIIME 2 which allowed us to perform quality control, data analysis, and create graphical and phylogenic visuals. With an overview of the microbiome, a literature review was done on abundant taxa and independent taxa were reviewed. Spotlight taxa were selected based on their potential impact on human health, climate cycles, and water taxa that are important globally for the quality of aquatic microbiomes. We found various Operational Taxon Units (OTUs) that are known to harbor antibiotic resistance such as bacteria in the families burkholderiales, pseudomonadaceae, sphingomondaceae, thermomicrobiaceae. We found various Operational Taxon Units (OTUs) that are known to play important roles in nutrient cycling, such as bacteria within the family rhodobacteraceae. We also concluded that there was a difference between OTUs found in rivers and creeks within Atlanta. This study showed that there are potential bacterial reservoirs for resistance in urban waterways.
Additional research and microbial ecology testing was then done on samples collected in Spring of 2022 to determine if any of the bacteria were resistant to antibiotics or carried tetracycline resistance or known pathogenic genes. The level of Escherichia coli (E.coli) found at the sampling sites was measured to check if it was within EPA federal guidelines. The E.coli was then isolated and tested to determine if the found strains are considered pathogenic or have antibiotic resistance.
Antibiotic-Resistant Bacteria (AR) Are bacteria that have gained opportunistic genes which allow them to not be affected by antibiotics that would normally be of a detriment to them.
-
School Work
- Antibiotic-Resistance in Atlanta's Urban Waterways Poster
Media
- Antibiotic-Resistant Bacteria in Atlanta's Urban Waterways Video
Powered by Acadiate
© 2011-2024, Acadiate Inc. or its affiliates · Privacy
