Your message has been sent.
 CLOSE SIDEBAR
CLOSE SIDEBAR
Fluorescence Image Analysis via Threshold Enhanced Alternative Morphology Guided Image Segmentation
4
student

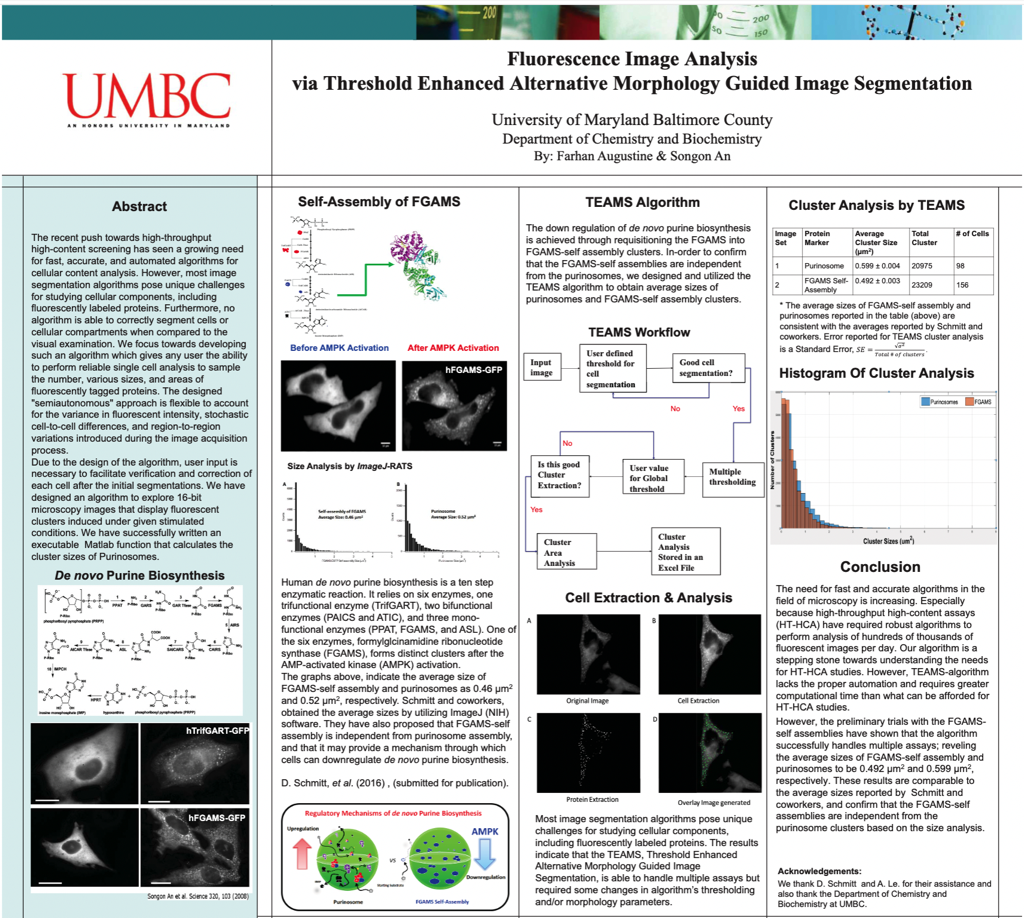
The recent push towards high-throughput high-content screening has seen a growing need for fast, accurate, and automated algorithms for cellular content analysis. However, most image segmentation algorithms pose unique challenges for studying cellular components, including fluorescently labeled proteins. Furthermore, no algorithm is able to correctly segment cells or cellular compartments when compared to the visual examination. We focus towards developing such an algorithm which gives any user the ability to perform reliable single cell analysis to sample the number, various sizes, and areas of fluorescently tagged proteins. The designed "semiautonomous" approach is flexible to account for the variance in fluorescent intensity, stochastic cell-to-cell differences, and region-to-region variations introduced during the image acquisition process.
Due to the design of the algorithm, user input is necessary to facilitate verification and correction of each cell after the initial segmentations. We have designed an algorithm to explore 16-bit microscopy images that display fluorescent clusters induced under given stimulated conditions. We have successfully written an executable Matlab function that calculates the cluster sizes of Purinosomes.
Resume / CV
- Resume / CV
Media
- Interview Dr. Tucker Harding,
- Scientific Poster 1 Interview
Powered by Acadiate
© 2011-2024, Acadiate Inc. or its affiliates · Privacy
